Covers & Featured Images

Postdoctoral Research Fellow · The Francis Crick Institute · London
I build computational tools to understand how Mycobacterium tuberculosis exploits host cell heterogeneity. My work sits at the intersection of live-cell time-lapse microscopy, deep learning image analysis, and quantitative cell biology — using single-cell tracking to ask why some infected macrophages clear bacteria while others succumb.
Characterising single-cell heterogeneity of M. tuberculosis infection in iPSC-derived human macrophages using high-throughput time-lapse microscopy. Deep learning segmentation and Bayesian tracking pipeline to trace bacterial burden dynamics at single-cell resolution across antitubercular drug treatments. Pipeline infrastructure built on Dask and OME-NGFF, designed for accessibility and integration with cloud-native, AI-ready approaches.
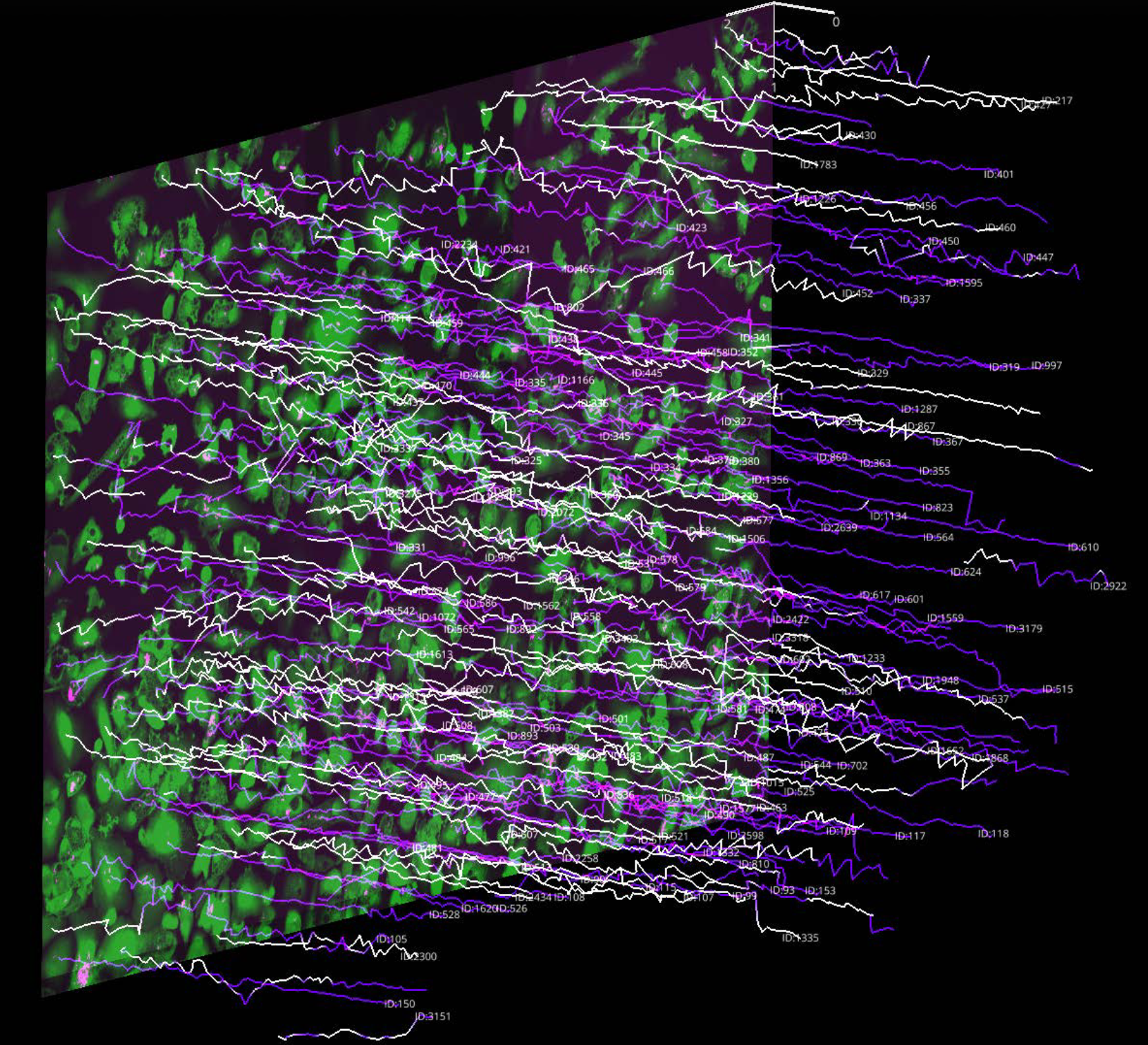
Reverse-engineering cell competition from single-cell observations using timelapse microscopy with deep learning segmentation and Bayesian tracking. Characterised mutant elimination dynamics in wild-type vs. ScrKD and RasV12 MDCK epithelium. This work was conducted on a bespoke automated epifluorescence microscope, the design and maintenance of which formed a core part of my doctoral training.
Implementing protein retention expansion microscopy (ProExM) on C. elegans touch receptor neurons.
Study of polarity-sensitive fluorescent dyes to probe membrane order in the T-cell immunological synapse, in conjunction with novel super-resolution image analysis techniques.
An investigation into the self-assembly of borazine-derivative monolayers using Metropolis Monte Carlo algorithms.


Mentored two teams during the annual Crick Data Challenge, guiding early-career researchers through Python-based image analysis and quantitative microscopy workflows.
Delivered lecture for postgraduate students on single-cell image analysis methods.
Helped design and deliver MSc-level courses in Computational and Systems Biology and Integrative Cell Biology, focusing on image analysis and machine learning methods.
Tutored mathematics, biology, chemistry, and physics from school child to graduate level, including private tuition and A-level classrooms.
Invited to present research on M. tuberculosis infection heterogeneity to visiting philanthropic donors.
Ran a public engagement stall at a philanthropy evening for prospective supporters of the Francis Crick Institute.
Hosted visiting school students at the Crick, introducing young people to life as a researcher and the science of infectious disease.
Participating in outreach at my former secondary school to encourage higher education progression — particularly meaningful as a first-generation university student.
Weekly volunteering throughout the pandemic and beyond, managing stock, coordinating large deliveries, and preparing food packages.